Theoretical ecology
The overall philosophy of the course is that in theoretical ecology and indeed in much of theoretical biology, there is no clear dividing line between biological problems and mathematical problems. Accordingly, the course covers both conceptual issues and methodological issues. The overall goal is to understand how a given set of assumptions leads to a particular model, and what each model predicts. Examples are drawn from basic models in ecology.
Co-taught with Greg Dwyer, 2014-2019, 2021-2023, 2025-, ECEV 42900
Pathogen evolution, selection, and immunity
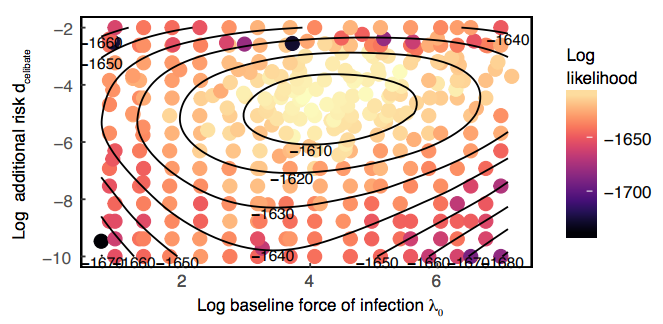
This module provides an introduction to modeling antigenically diverse pathogen populations. The first part of the course will introduce multistrain compartmental models and potential mechanisms of competition. These simple models will be contrasted with models with more complex assumptions (e.g., multiple forms of immunity and spatial structure). We will review how to statistically fit multistrain models to longitudinal data from individuals and time series data from populations. The second part of the course will show how, using the coalescent as a neutral expectation, evolutionary pressures can be quantified using sequence data. We will detail bioinformatic methods to build phylogenies, quantify selective pressures and estimate pathogen population structure. Methods to measure pathogen phenotypic similarity and antigenic evolution, such as antigenic cartography, will be introduced.
Co-taught at SISMID with Trevor Bedford, 2015-2019, 2021-2023 slides

Evolutionary and genomic medicine
Evolution is regularly investigated in free-living organisms, but some of its most fascinating and important examples occur in the interface between free-living and non-free-living states. In this course, we use evolutionary and ecological principles to study the dynamics of clonal populations in multi-cellular organisms relevant to human medicine. The emphasis is on the evolution of pathogens, the evolution of cells of the immune system in response to pathogen invasion, and the population genetics of cancerous cells in light of recent cancer genomic studies. This is a mixed lecture and seminar-style course with substantial reading, presentation, and discussion by students.
Co-taught with Chung-I Wu, 2015-2019, BIOS 23365/ECEV 33365

Defensive programming in R
A very short course in how to prevent and isolate bugs and stay calm when programming with RStudio.
Presented at the BSD Quantitative Bootcamp for incoming graduate students in 2017, complete materials